AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog

import plotly.express as pxĪ = np.repeat(np.arange(1, 10), 4, axis=-1)ĭf = pd.DataFrame(a, columns=)ĭf.category = Ĭolor_discrete_map=, the yellow points having different levels of opacity, and the blue points having different levels of opacity. To expand slightly on the example you gave, what I am effectively after is the following, with the color being set by the category (in my cases, diagnosis) and each category having different levels of opacity within it. Or, you can set the marker attributes for each point, but they then become their own trace, and you can’t group them in the legend. I am effectively after another axis of information within the plot.Īs I’m reading things, you can’t have points in the plot that are part of the same trace having different opacity - you can only set marker attributes for an entire trace. 
I think I’m leaning towards either Plotly can’t do the thing I want it to do, or I’m thinking entirely the wrong way about it. That almost works, but you still end up with a different category label for each point. Which I can see being a useful things for some scenarios I guess, but not this one.Īny ideas on how I would go about doing this, would be much appreciated. The way that I initially tried, which makes the most sense to me, would have been to pass a column from the dataframe to the opacity variable when I first made the plot, but opacity accepts only a single value. It gives me variable transparency on a per sample basis, but it also results in multiple traces, one for each sample, which means they are then not color coded or grouped in the plot and legend, Which is less than ideal. SampleId, patientId, diagnosis, cancerType = row]įig.add_trace(go.Scatter3d(mode='markers',Īnd this sort of works. 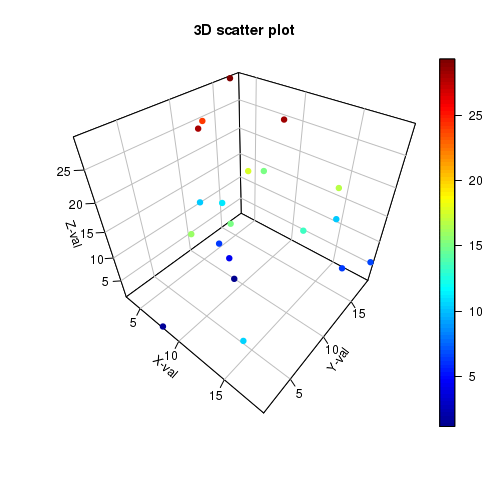
The other way that I tried was to construct the figure from graph objects like this: fig = go.Figure()Ĭolors = px.Ĭolor_mapping = dict(zip(categories, colors))ĭf = df.map(color_mapping) Doing it this way, I could set an opacity for the markers of each diagnosis, but I don’t think I could do it for each sample. The traces each contain multiple samples, and I’d like to change the opacity on a per sample basis. Hover_data=df],īut then when I iterate over the traces in the figure like this: for trace in fig.data: I can create a plot in plotly express like this: fig = px.scatter_3d(df, x="x", y="y", z="z", I have a dataframe with columns for x,y,z, sample_id, patient_id, diagnosis and sample_purity. I am trying to create a 3d scatter plot with the opacity of each point drawn from a column in my dataframe.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
